Microsatellites Explorer Help Page
This is the homepage of Microsatellites Explorer. It is comprised of high level metadata about the
reasearch
done, mainly statistics by biological domain , by repeating sequence and biological species:
From here you can navigate to other pages using the navigation bar , or you can
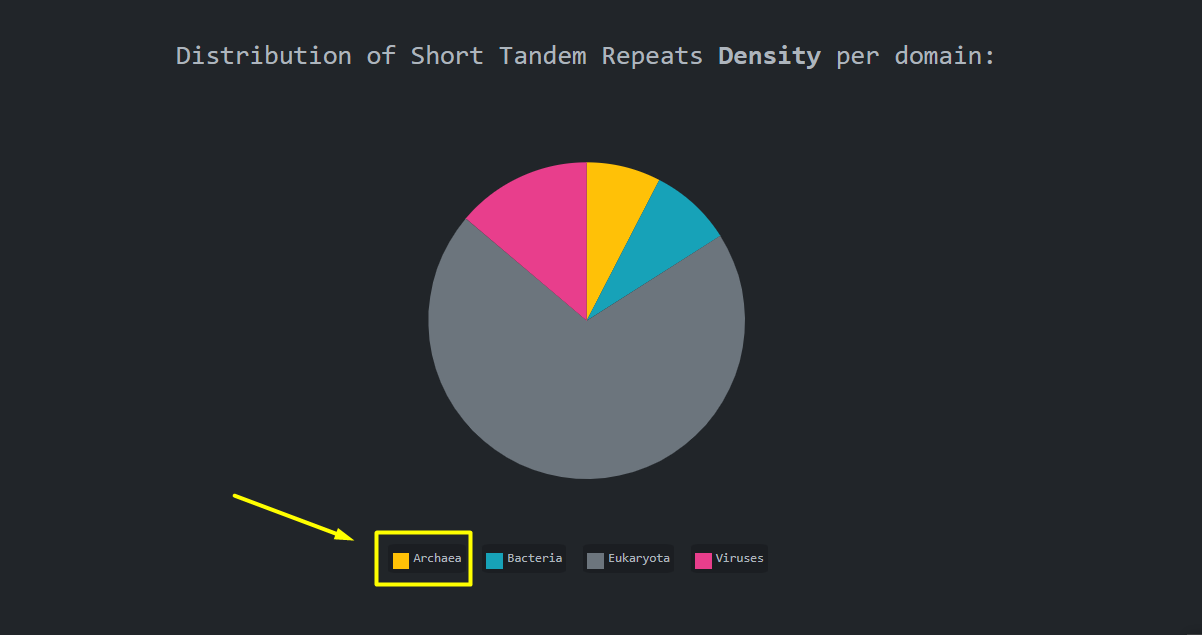
navigate to the statistics per domain using the pie chart legend items
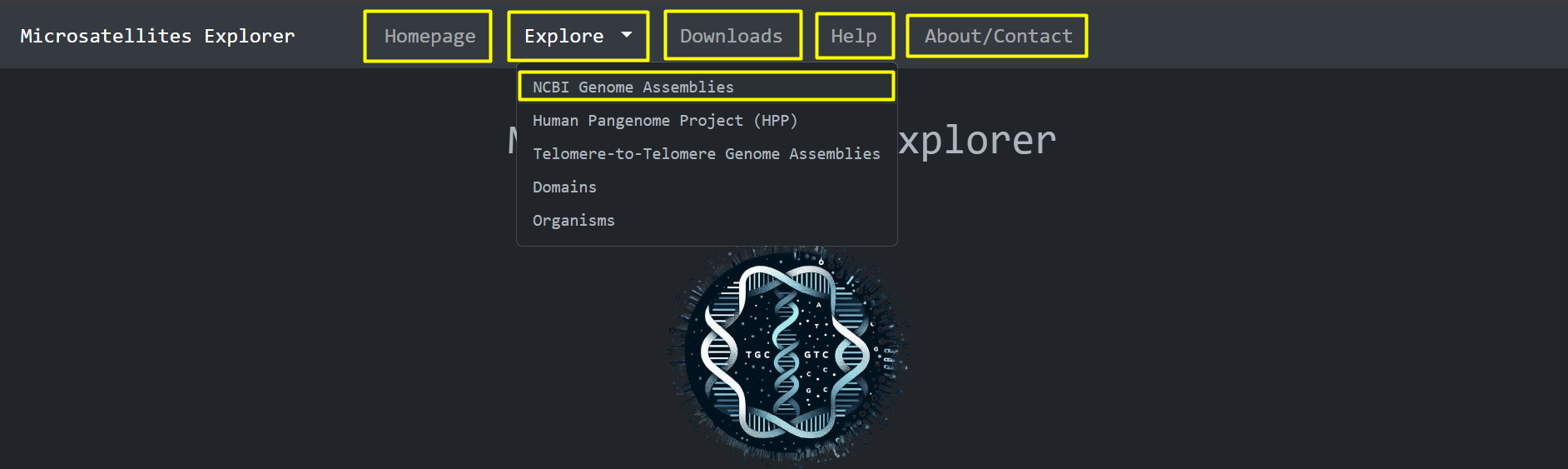
Navigation Bar:
Legend Items:
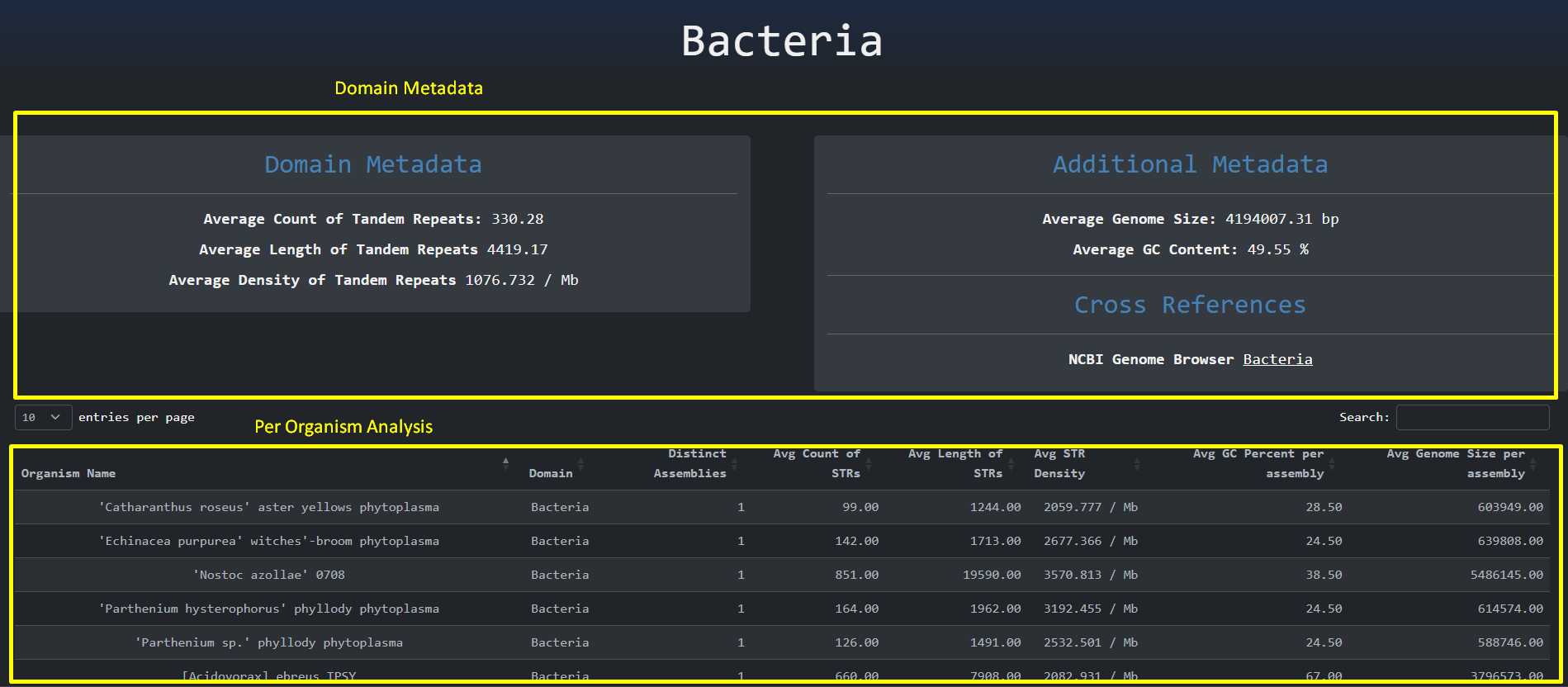
By clicking the legend items we are presented with the statistics per biological domain page. As the
name implies, this page
is used to perform a statistical analysis on the most relevant properties per domain and a further
analysis per biological organism.
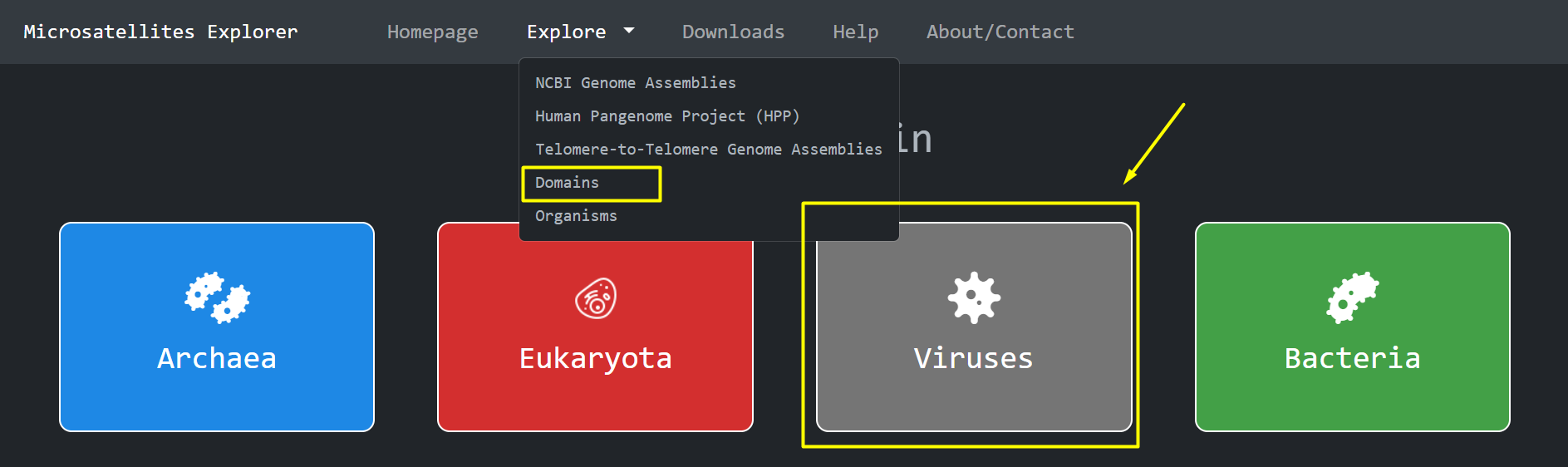
Another way to access the different domains is through the Navigation Bar > Explore Domains
The domain metadata
are organized at the upper part of the page whereas the organismal analysis is presented as a table on
the lower part.
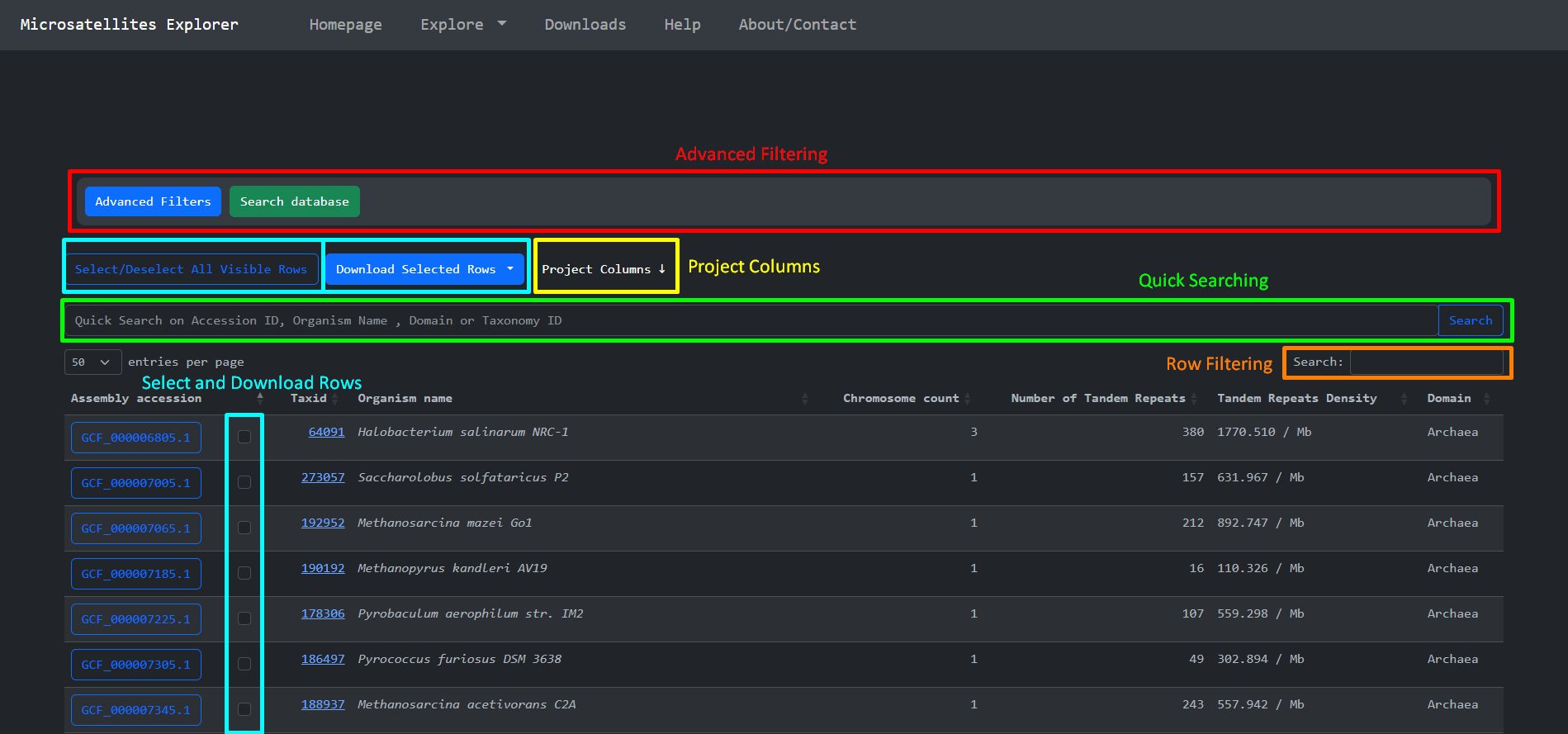
By clicking the Explore item at the navigation bar we are presented with a page that is used to
effectively search , access and download parts of the
Short Tandem Repeats dataset we produced. This page consists of some features that we implemented to
explore the dataset. These are:
-
Advanced Filtering
-
Quick Searching
-
Select and Download
-
Project Columns
-
Row filtering
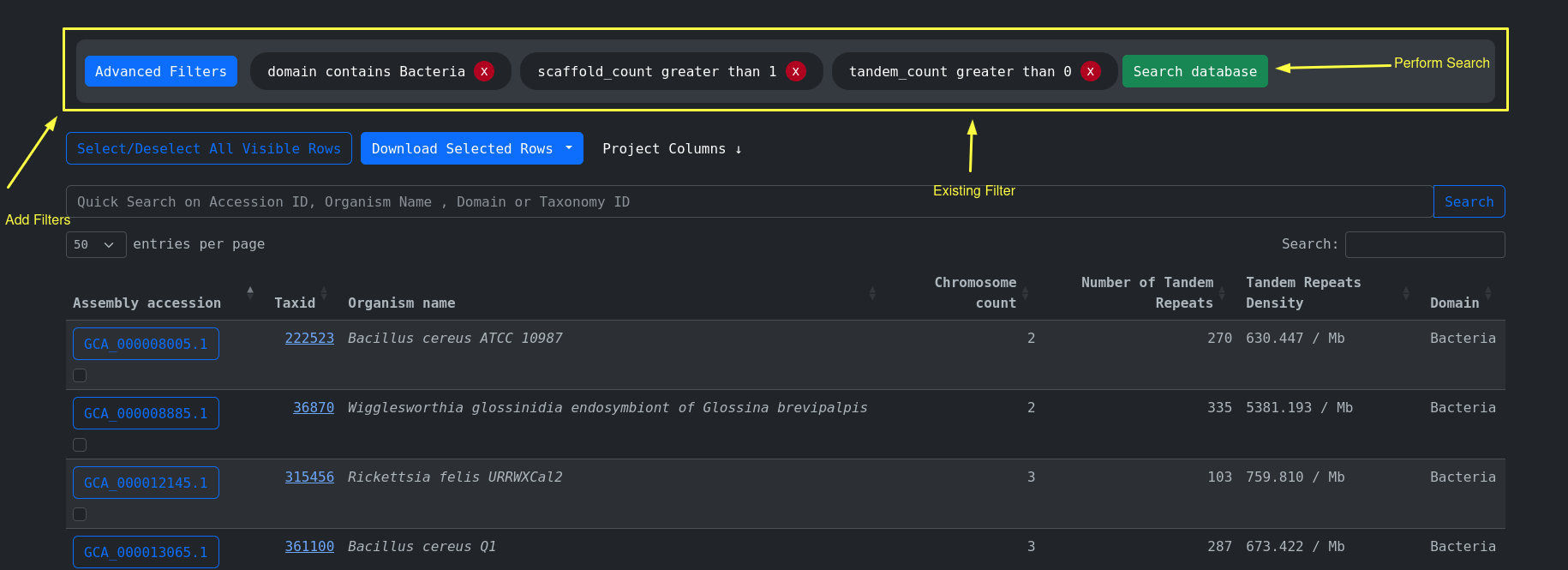
Advanced filtering is used to perform rich querying using all the available columns present on the accession
metadata. For example find every accesson that
comes from the domain of Bacteria and has more than 1 Chromosome and short Tandem Repeats present:
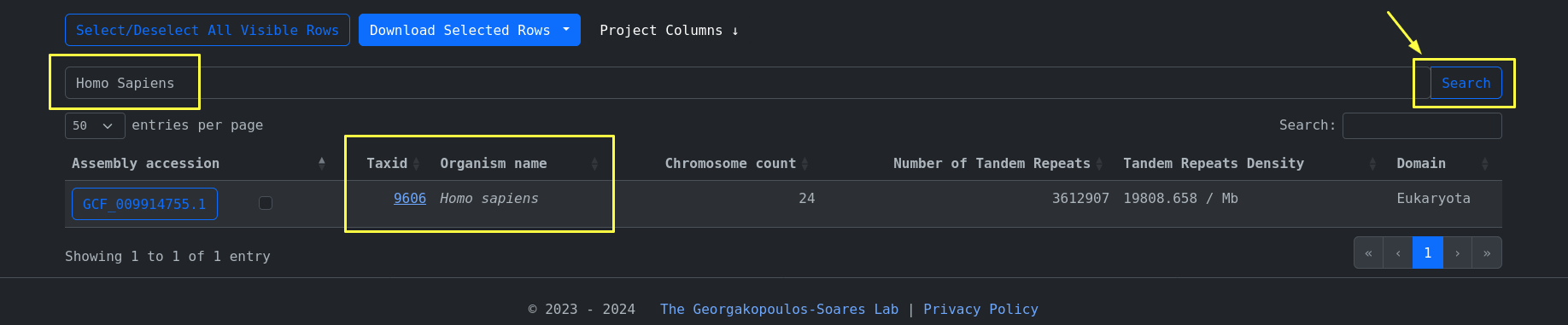
The quick search feature is used to search on the most frequently used columns, for example:
organism_name=Homo Sapiens
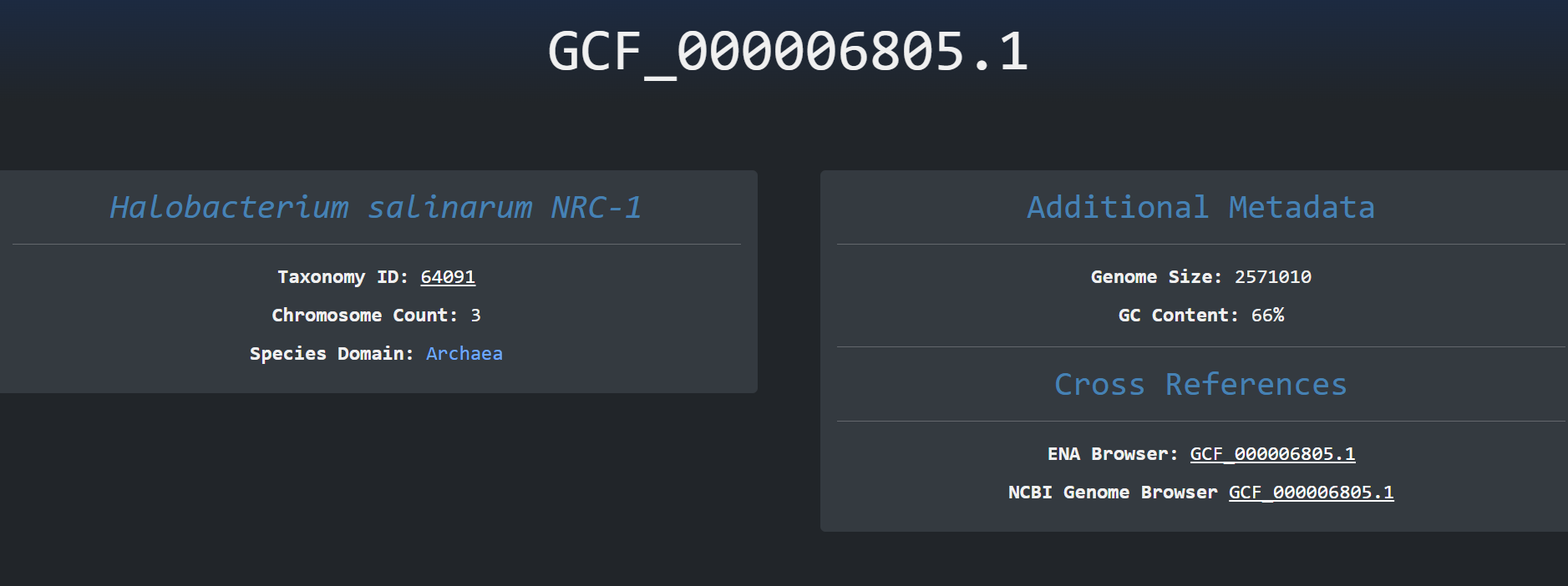
We can then click on the accession id button of a specific row and navigate to the tandem repeats page of
this specific accession. Here we can
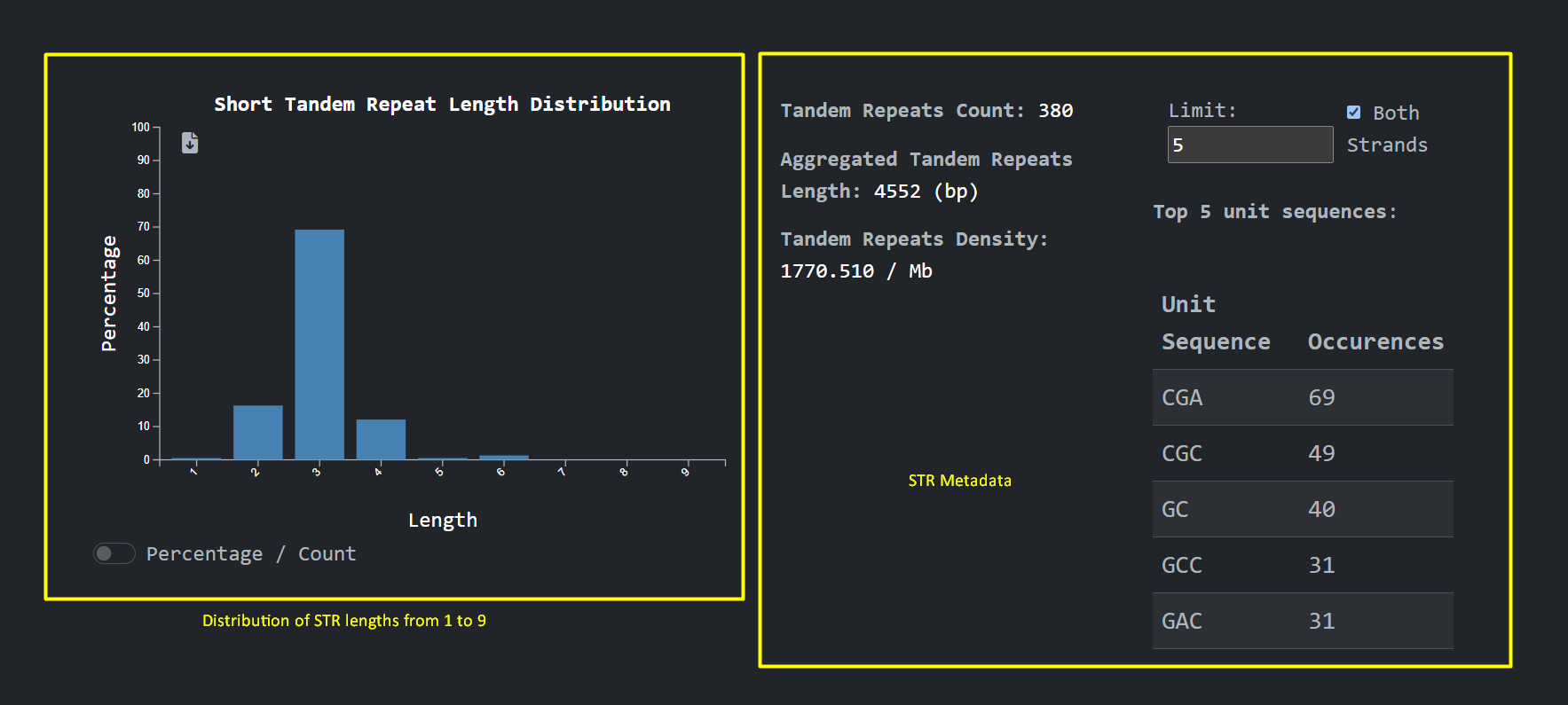
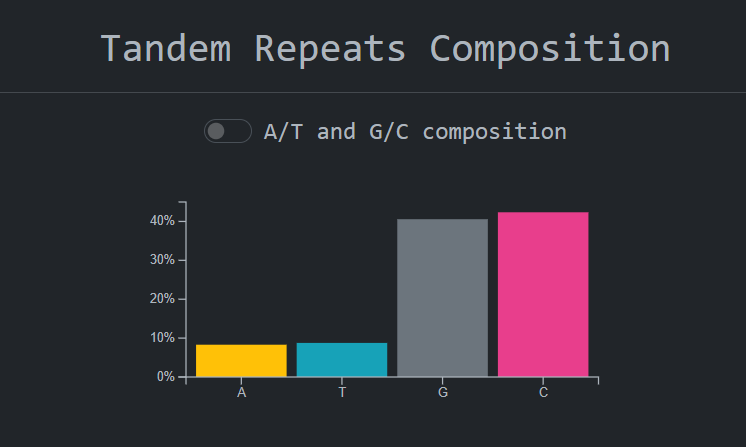
see calculations on tandem repeats , accession metadata, graphs about the distribution on tandem repeats
units lengths and cross references to
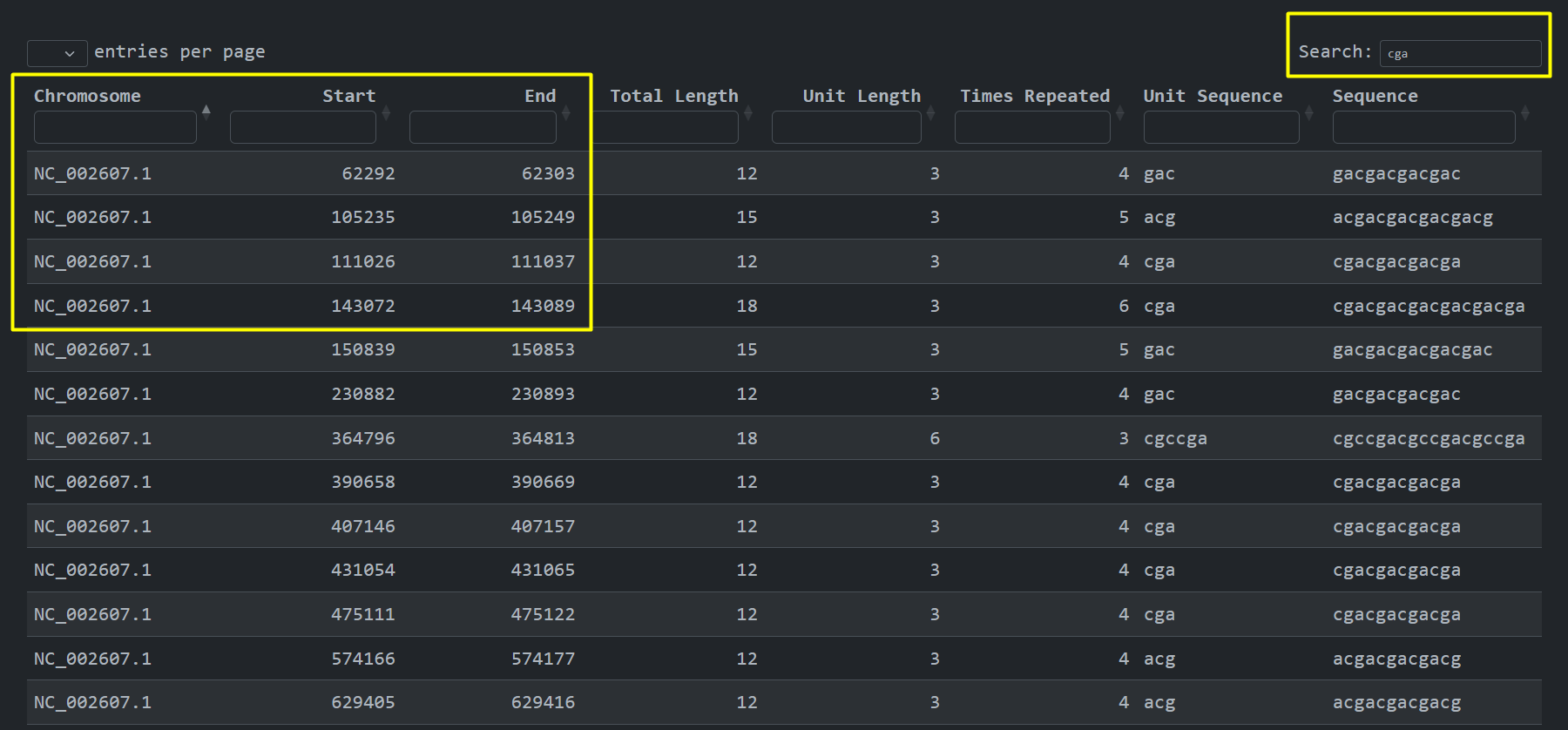
other public sources. We can also see where the short tandem repeats are present at the genome , Chromosome
, Start , End:
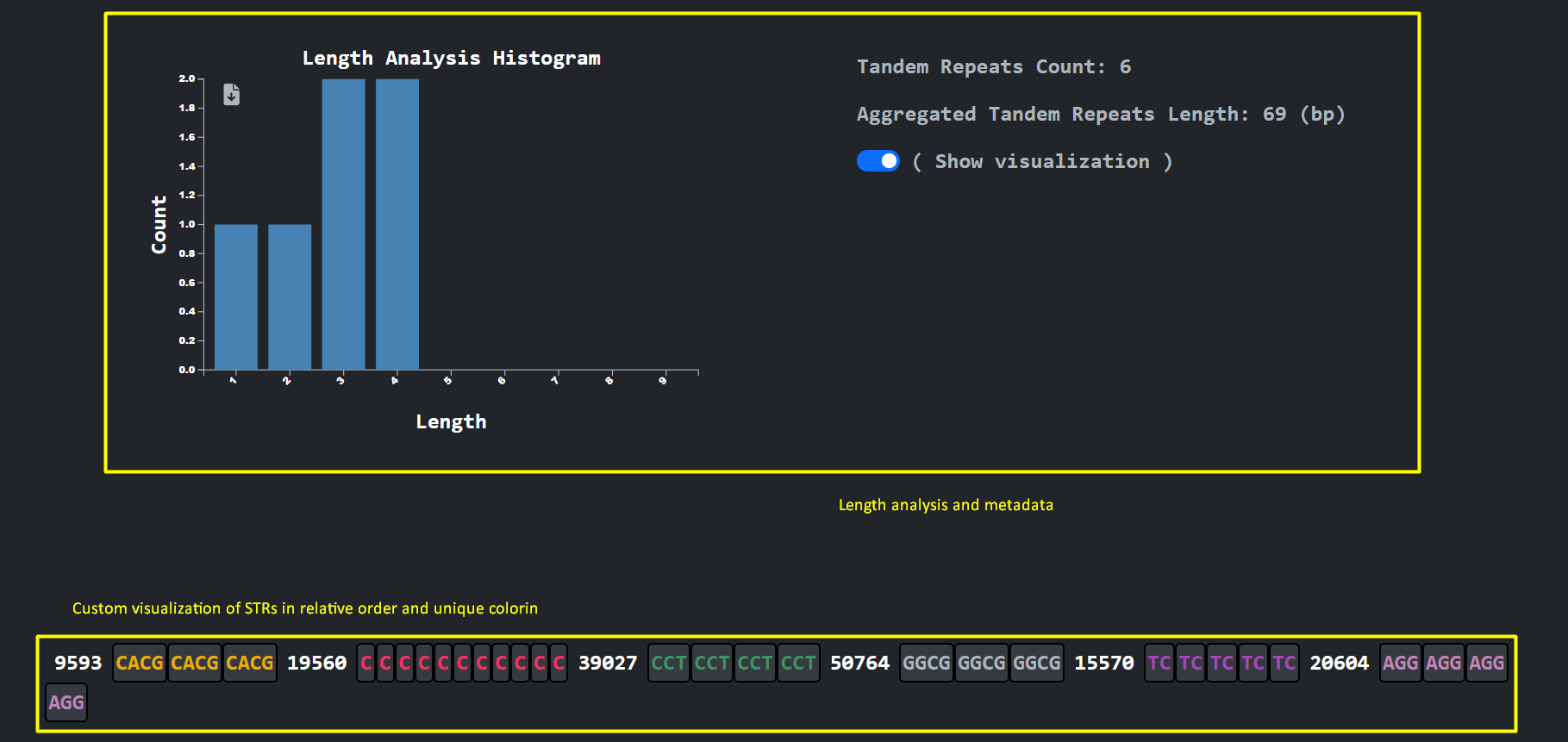
An analysis and visualization per chromosome that the user selects provides the user with the ability to inspect the different STR unit lengths along with an arbitrary presentation of the unit sequences using unique coloring and a magnitude number to denote the gap between consecutive STR sequences
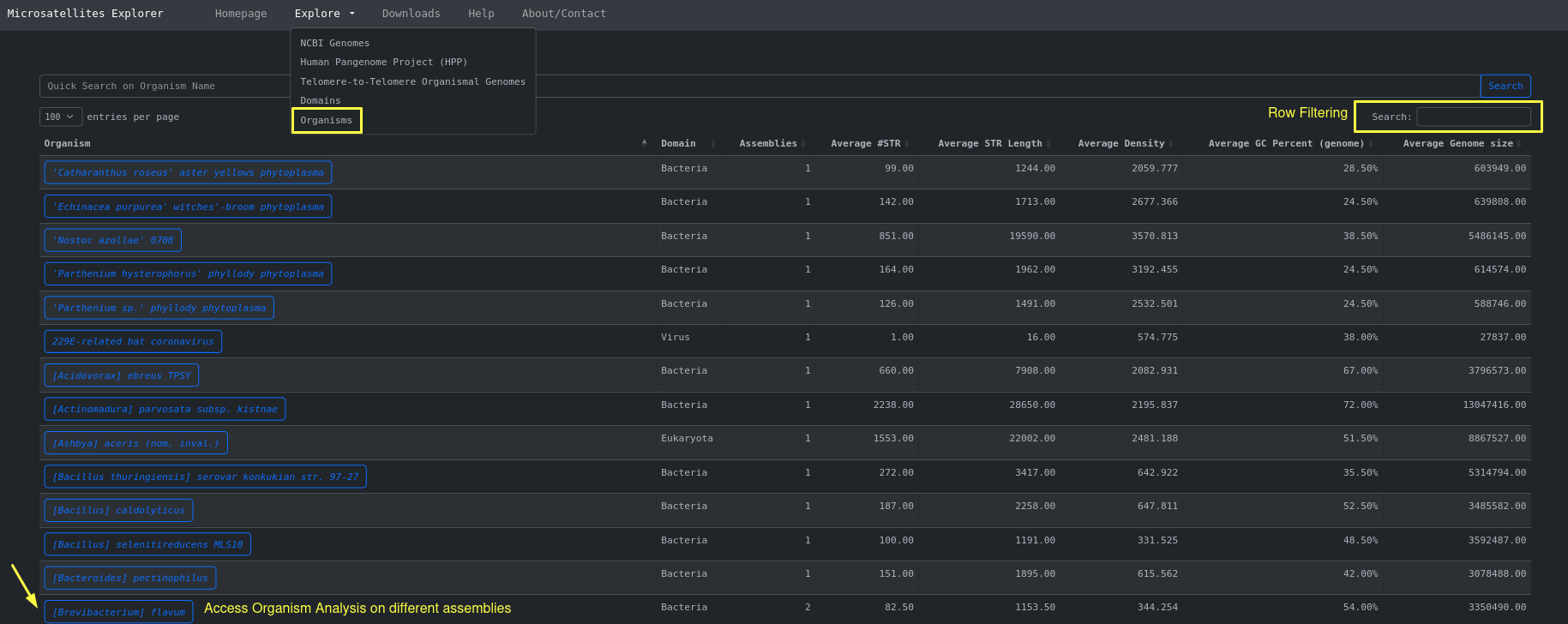
Another view of the same data ( NCBI organismal genomes ) is an aggregation of the different assemblies based on Organism Name. This can be
accessed through Explore > Organisms
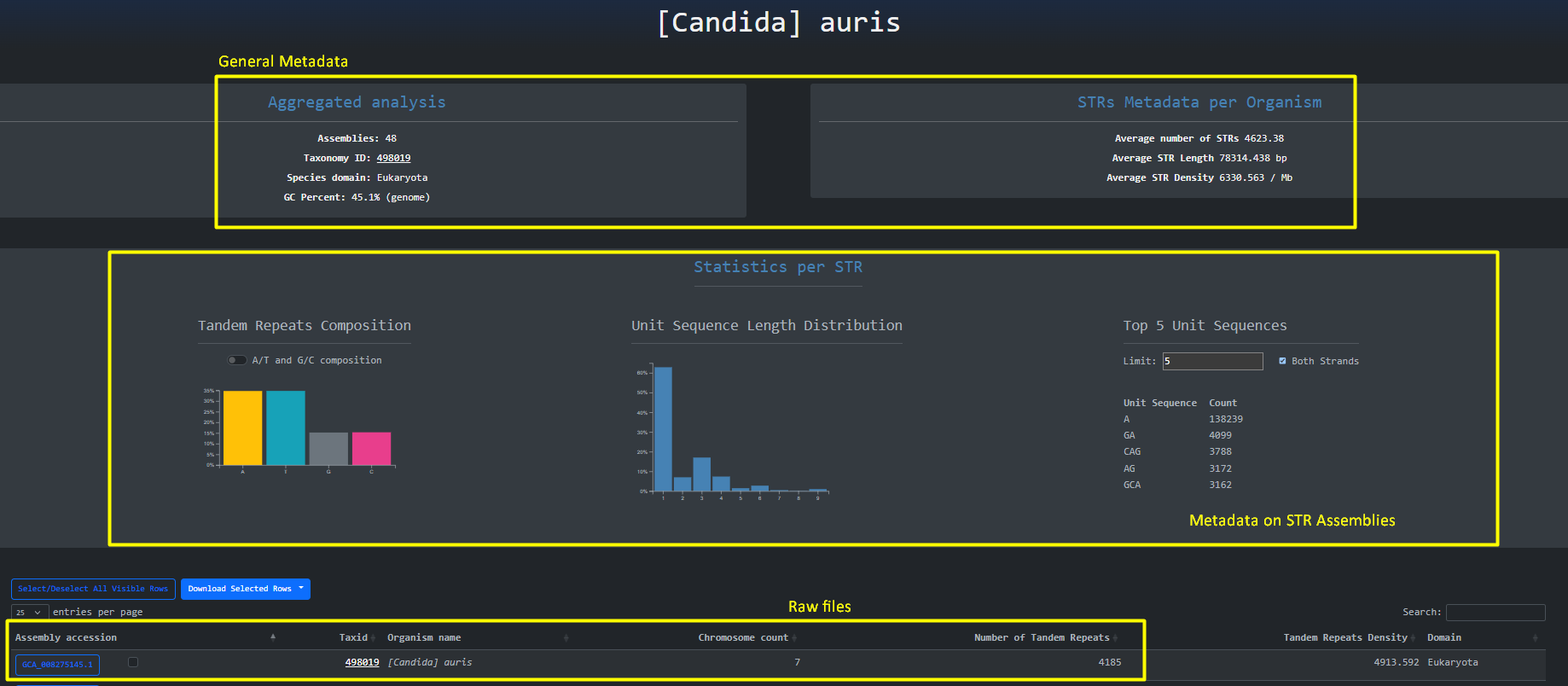
By clicking the organism name button we are being redirected to the aggregated analysis done on the organism for the different assemblies. Here we
can gain a high level overview about STRs across assemblies and Inspect Filter and Download the raw files.
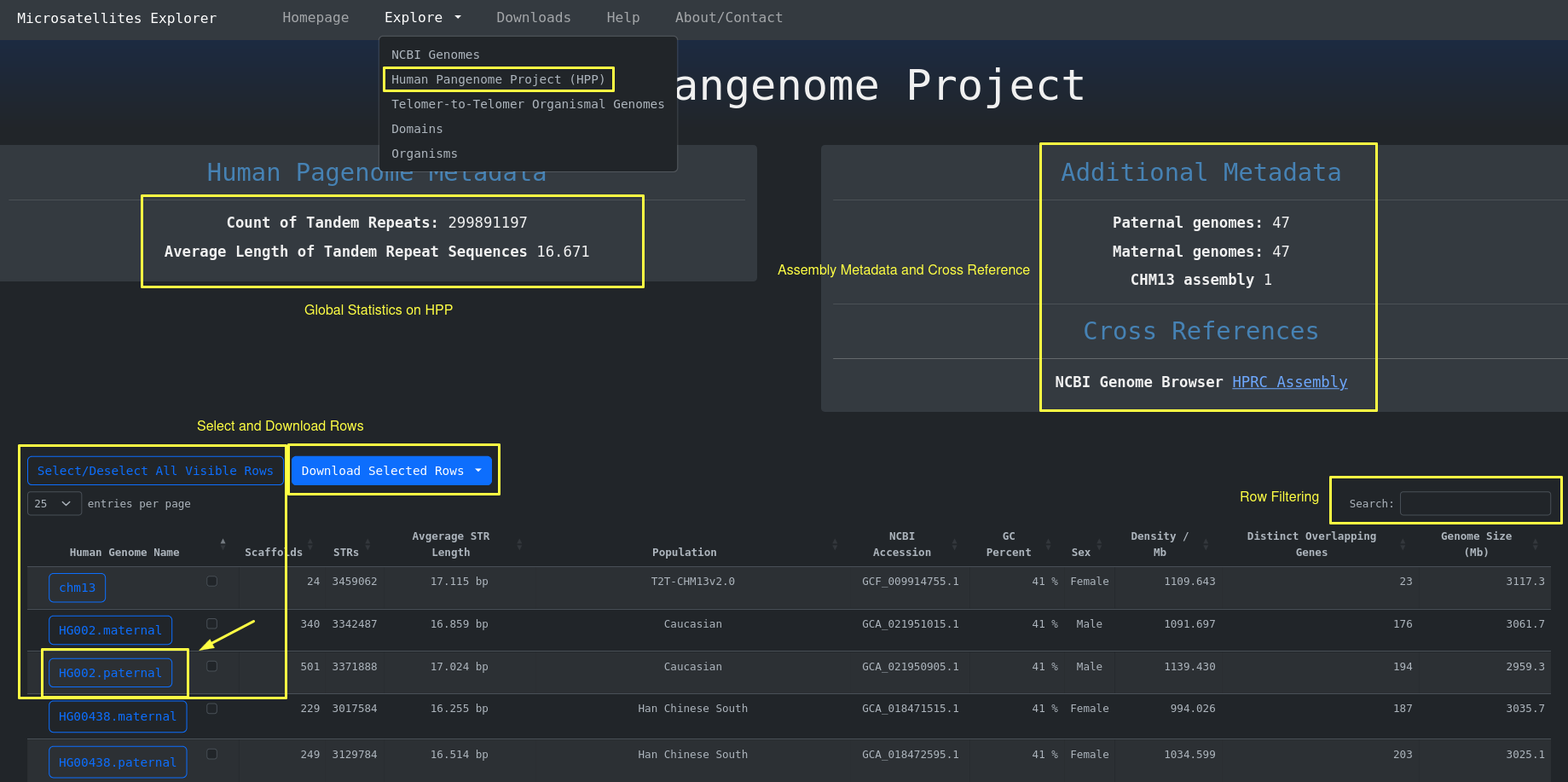
In order to access the Human Pangenome Table with the 95 human genomes including their STR metadata you go
to
Explore > Human Pangenome Project (HPP)
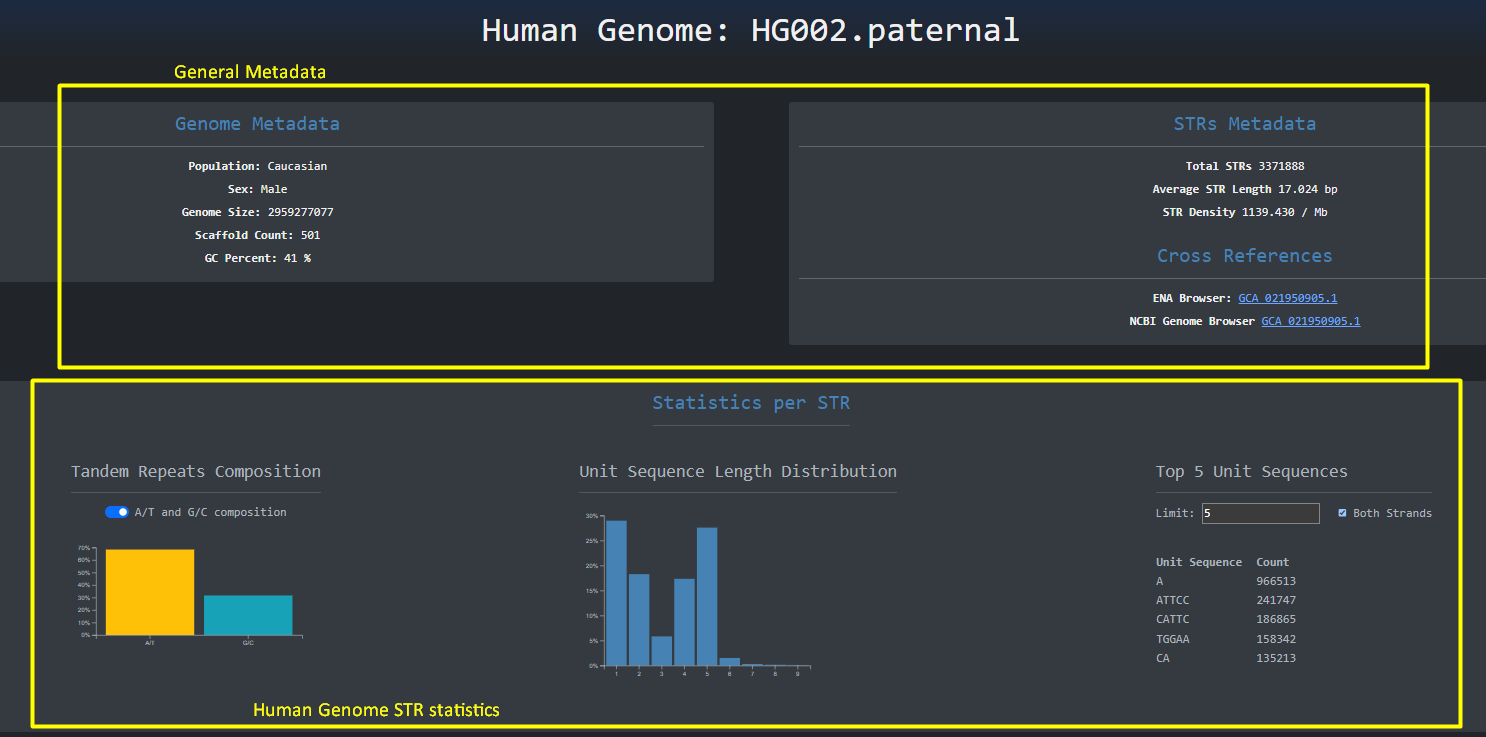
By clicking the button with the human genome on the left side of the table we are redirected to a page
similar to the NCBI genome assemblies where the full human genome is analyzed in the context of short tandem
repeats. Here
as the headings imply we have some general metadata and some metadata about STRs.
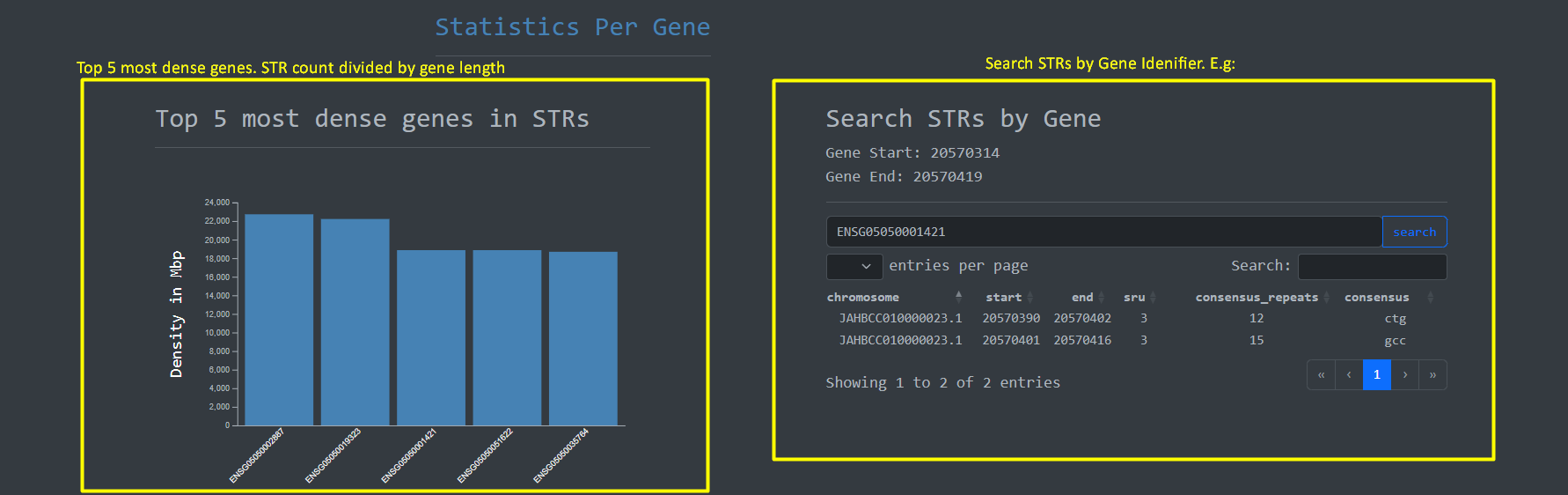
Given the importance of genes on the human genome we implemented a feature to perform Short Tandem Repeat
searches on genes based on Gene ID along with
a bar plot for the top 5 most dense genes in terms STRs :
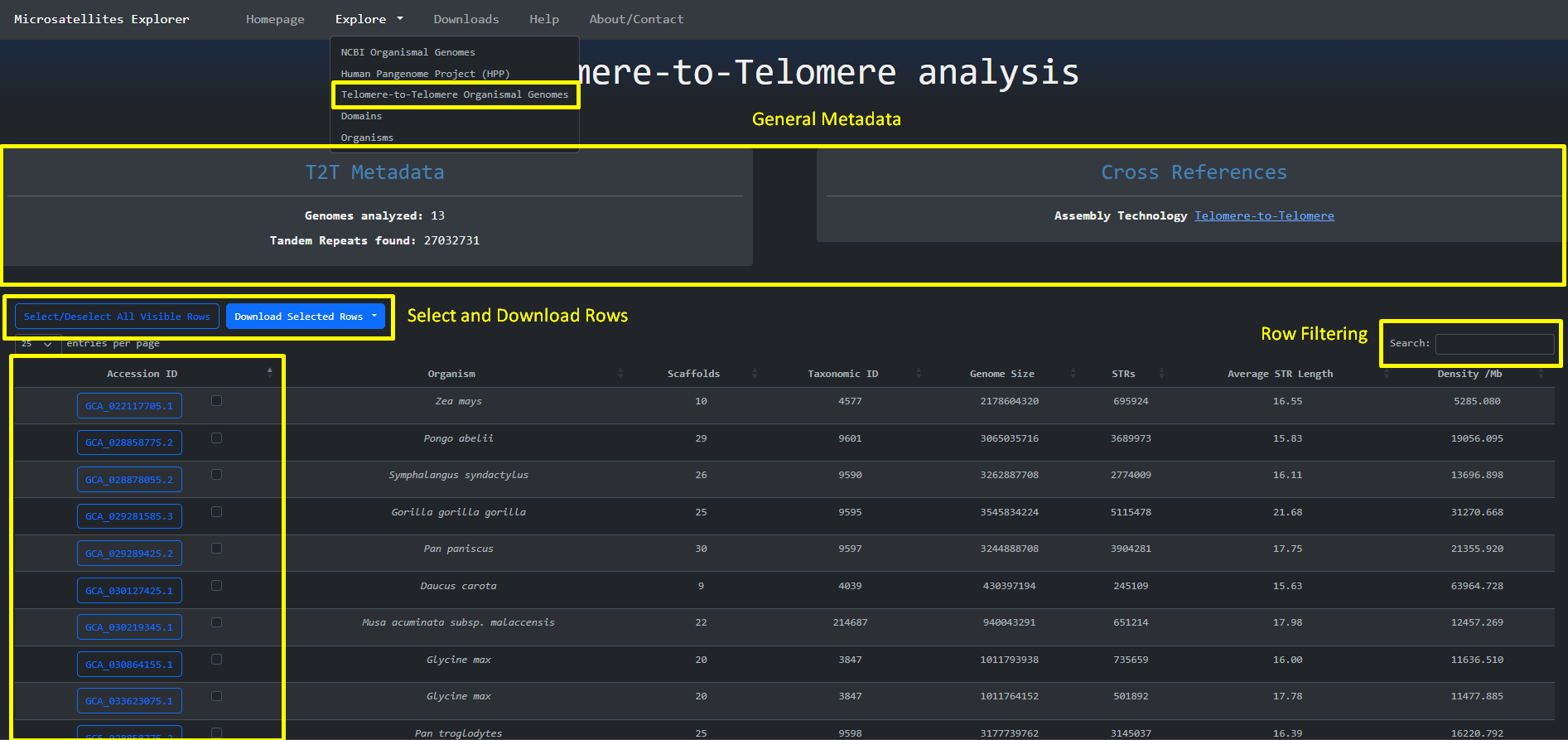
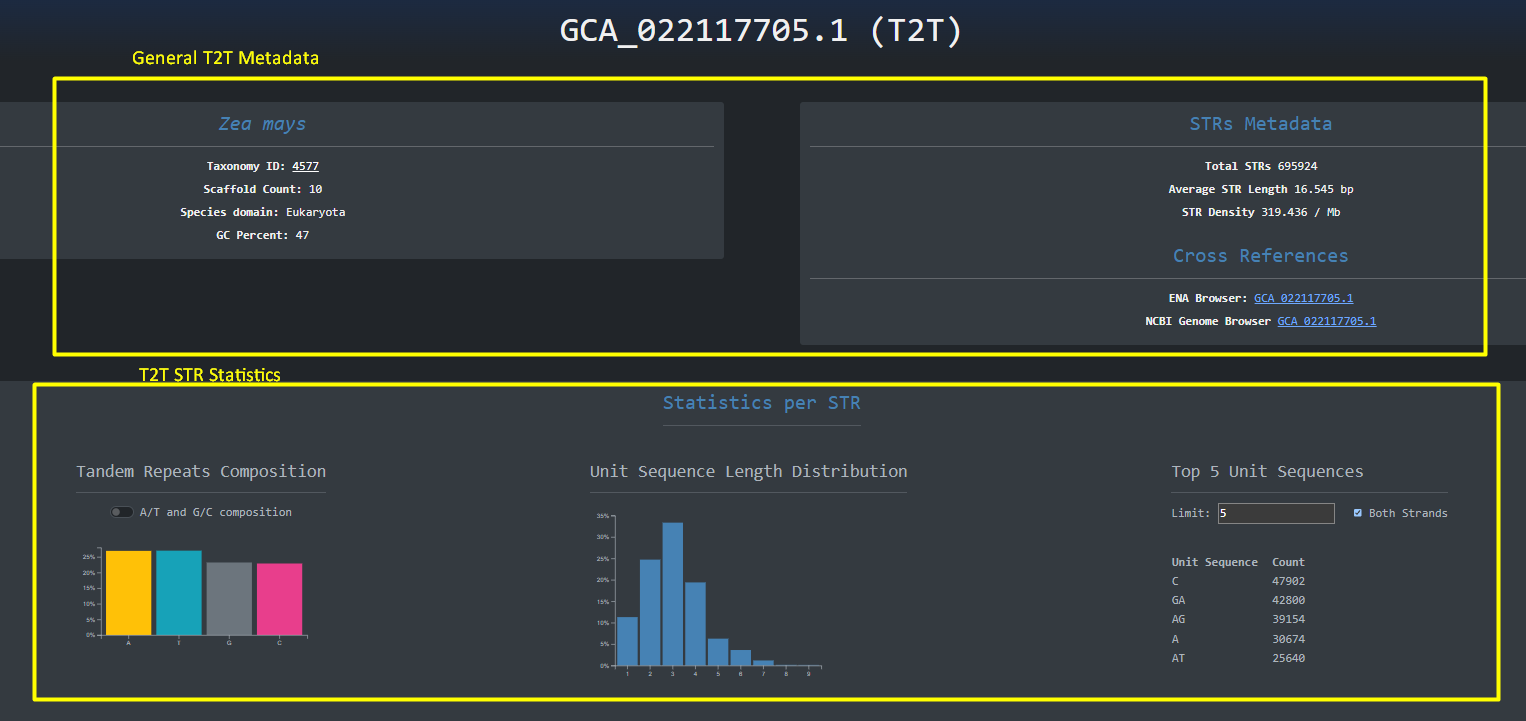
The same design principle is followed for the Telomere-to-Telomere genomes that are accessed through Explore > Telomere-to-Telomere Organismal Genomes:
We can then navigate to the specific assembly by clicking the blue button with the accession_id on the left
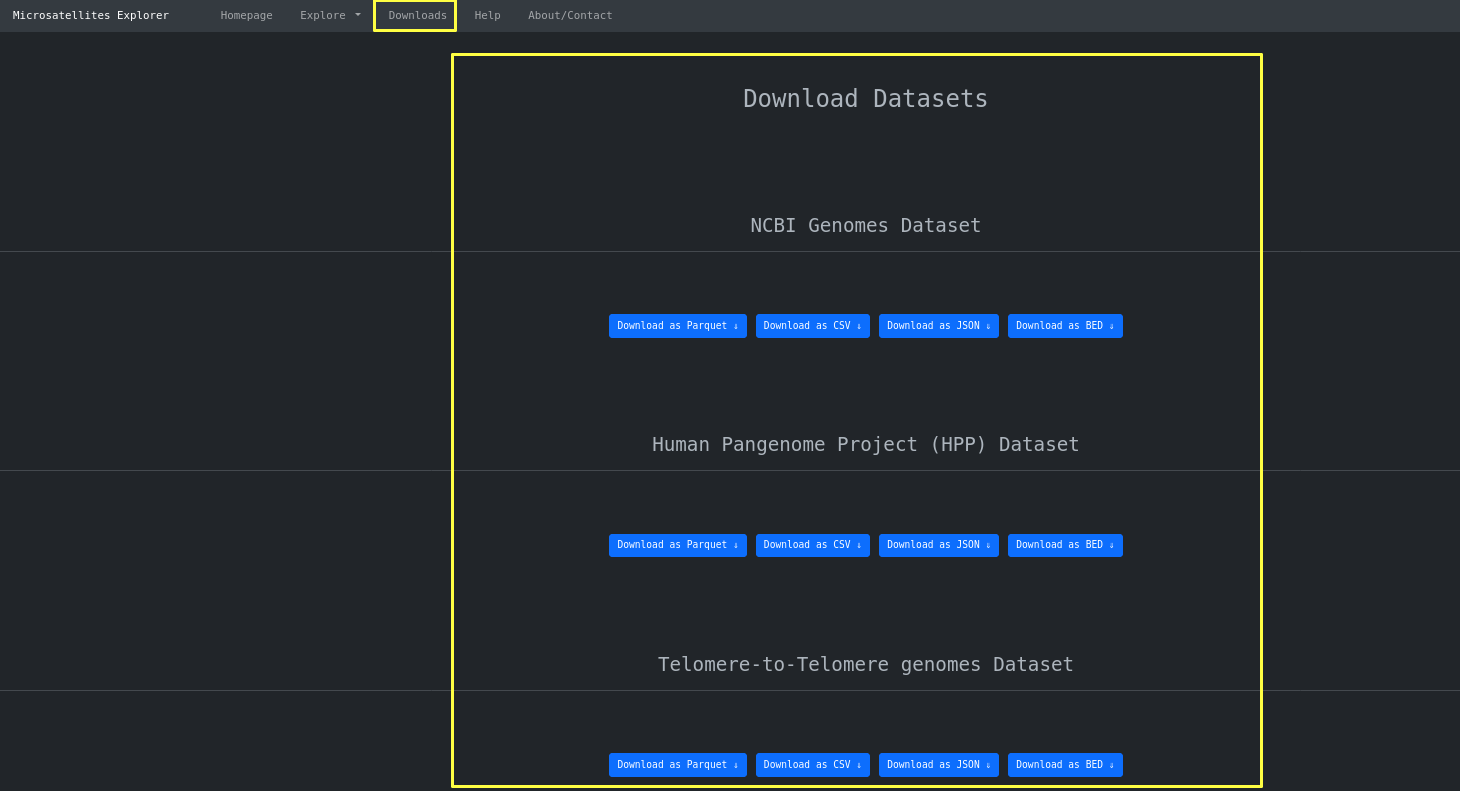
Finally you can use the downloads page to download the dataset as a whole in three different formats
depending on the use. For example CSV for
spreadsheet use, JSON for app development and Parquet.snappy for HPC use.